
In view of the current dramatic events in Afghanistan many wonder why the extensive international efforts to bring some stability to the country have failed so miserably.
In this post, we will present and analytically examine a fascinating theory that seems to be able to explain political (in-)stability almost mono-causally, so read on!
Continue reading “The “Youth Bulge” of Afghanistan: The Hidden Force behind Political Instability”



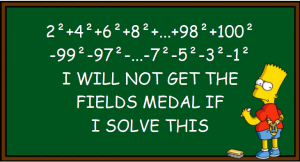 This time we want to solve the following simple task with R: Take the numbers 1 to 100, square them, and add all the even numbers while subtracting the odd ones!
This time we want to solve the following simple task with R: Take the numbers 1 to 100, square them, and add all the even numbers while subtracting the odd ones!


